It took twenty years to fully work up the X chromosome. This now became possible with nano pore sieves to sort out the complex portions not addressed.
The big news is that we can now tackle the other 23 chromosomes within a year.
This will be s huge accomplishment and set the stage for a huge expansion of the science..
Human X chromosome completely sequenced for the first time
July 14, 2020
The human X chromosome has been completely sequenced from end to end for the first time
As close as we’ve gotten to sequencing the entire human genome, there are still a few gaps. But now geneticists have plugged a major one in a landmark new study, by sequencing the entire human X chromosome from end to end, covering more than three million base pairs that were previously unmapped.
The Human Genome Project was one of the most ambitious scientific undertakings of all time. Between 1990 and 2003, an international team of scientists worked to sequence the human genome in high detail, with the end result being an almost complete blueprint for the human species.
But the emphasis there is on “almost.” The first version covered a little over 92 percent of the human genome, with more than 99.99 percent accuracy. Later revisions closed some of the gaps, but others still remain.
The biggest gaps were located at the center and ends of chromosomes, which are known as centromeres and telomeres respectively. These regions are categorized by huge sections of repeating sequences, which can be hard to sort out.
Now an international team of geneticists has patched these gaps up. The researchers managed to sequence the entire X chromosome for the first time, from telomere to telomere.
In humans, the X chromosome is one of the sex-determining chromosomes passed down from parent to child. Generally, a zygote that receives two X chromosomes – one from each parent – will be usually biologically female, while an X and a Y chromosome results in a male.
In this case, the team didn’t sequence an X chromosome from a normal human cell. Instead, they studied a special type of cell that contains two identical chromosomes, part of a model genome dubbed CHM13.
One problem with sequencing genomes is that the technology traditionally can only read short segments of DNA at a time, leaving scientists to piece it all together. Unfortunately, that can be particularly difficult when those segments repeat over and over, which the team likens to assembling a jigsaw puzzle that’s all one color.
To finish the puzzle, the team used new techniques that read much longer sequences at a time. One of these is what’s known as “nanopore technology,” which funnels single molecules of DNA through a tiny hole and sequences them by detecting changes in the current flow.
“These repeat-rich sequences were once deemed intractable, but now we’ve made leaps and bounds in sequencing technology,” says Karen Miga, lead researcher on the project. “With nanopore sequencing, we get ultra-long reads of hundreds of thousands of base pairs that can span an entire repeat region, so that bypasses some of the challenges.”
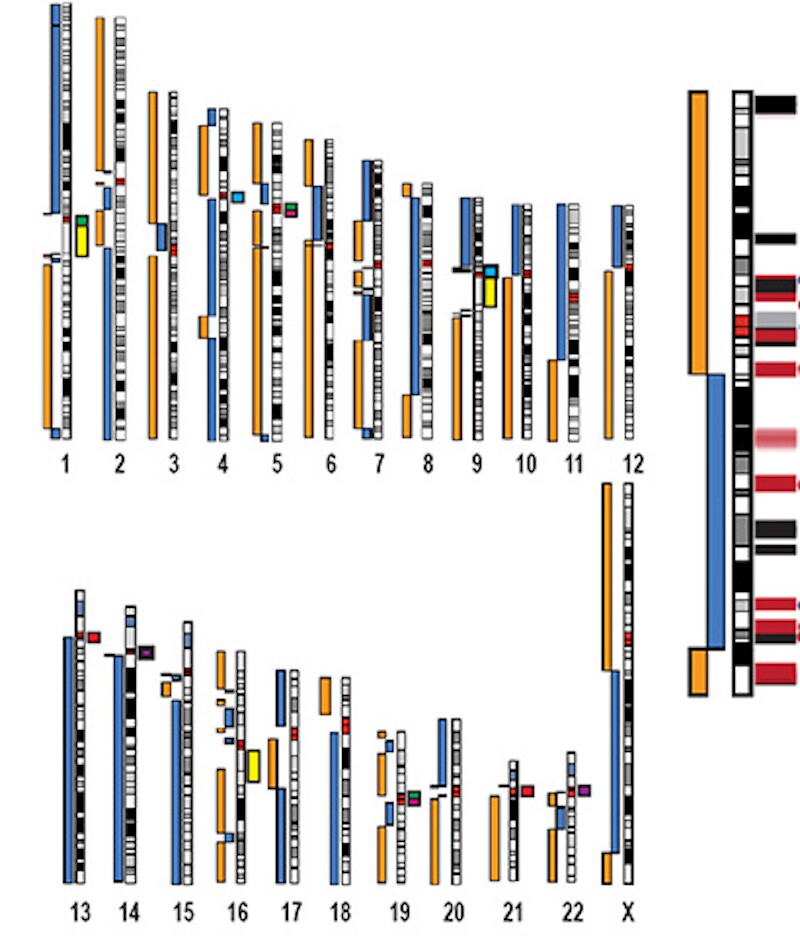
Ideograms showing the completeness of chromosomes in the human CHM13 genome. Completeness is measured in terms of the blue and orange rectangles to the left of each chart. The X chromosome is enlarged on the right
Miga et al., Nature 2020
Using this technique, the team was able to fill in a huge gap in the centromere, comprising some 3.1 million base pairs of repetitive DNA. And these previously unexplored areas may prove to be particularly valuable to science.
“We’re starting to find that some of these regions where there were gaps in the reference sequence are actually among the richest for variation in human populations, so we’ve been missing a lot of information that could be important to understanding human biology and disease,” says Miga.
But the X chromosome is just the beginning. There are 23 other chromosomes left, and the project plans to map them all by the end of 2020. In doing so, we may soon have a complete human genome sequence.
The research was published in the journal Nature.